This tutorial demonstrates how to access and visualize NASA CASALS (Concurrent Artificially-intelligent Spectrometry and Adaptive Lidar System) L1B geolocated waveform data from AWS S3.
About CASALS L1B Data¶
Data Organization:
Pulse: A single laser ‘shot’ with unique timestamp and Tx/Rx waveforms (up to 256 possible wavelengths/channels)
Track: A collection of pulses with the same wavelength, illuminates surface in along-track direction (direction of flight)
Sweep: A collection of coincident pulses spanning available wavelengths, illuminates surface in across-track direction
File Naming Convention: casals_LEVEL_UTC_VERSIONID_REV.FORMAT
Example:
casals_l1b_20241118T165452_001_02.h5LEVEL: Product level (l1a, l1b, l2a...)
UTC: J2000 UTC start time (yyyymmddThhmmss)
VERSIONID: Version number (001, 002...)
REV: Revision number (01, 02...)
FORMAT: File format (.h5)
File Sizes (per ~20-30 seconds of data):
| File Type | Size | Notes |
|---|---|---|
| L0 (TDMS) | 19 GB | Native LabView |
| L1A (H5) | 13 GB | Internal compression |
| ARD (H5) | 100 MB | No waveforms saves space |
| L1B (H5) | 14 GB | Contains reference height and waveforms |
import xarray as xr
import matplotlib.pyplot as plt
import numpy as np
import boto3
from botocore import UNSIGNED
from botocore.config import Config
import s3fs1. List available data files¶
First, let’s see what L1B files are available in the S3 bucket. We are only looking at the first 10 files and showing only hdf5.
# Configure S3 client for anonymous (public) access
s3 = boto3.client('s3',
region_name='us-west-2',
config=Config(signature_version=UNSIGNED))
# Bucket information
bucket_name = 'casals-data'
prefix = 'lidar/'
# List files
print("Available CASALS data files:\n")
response = s3.list_objects_v2(Bucket=bucket_name, Prefix=prefix, MaxKeys=10)
if 'Contents' in response:
for obj in response['Contents']:
# Only show H5 files
if obj['Key'].endswith('.h5'):
print(f"{obj['Key']}")
print(f" Size: {obj['Size'] / (1024**3):.2f} GB")
print(f" Last Modified: {obj['LastModified']}\n")
else:
print("No files found")Available CASALS data files:
lidar/2024-11-12/casals_l1b_20241112T163923_001_02.h5
Size: 3.04 GB
Last Modified: 2025-12-11 15:02:45+00:00
lidar/2024-11-12/casals_l1b_20241112T163934_001_02.h5
Size: 1.30 GB
Last Modified: 2025-12-11 15:02:45+00:00
lidar/2024-11-12/casals_l1b_20241112T163941_001_02.h5
Size: 1.57 GB
Last Modified: 2025-12-11 15:02:45+00:00
lidar/2024-11-12/casals_l1b_20241112T164250_001_02.h5
Size: 12.98 GB
Last Modified: 2025-12-11 15:09:39+00:00
lidar/2024-11-12/casals_l1b_20241112T164315_001_02.h5
Size: 12.95 GB
Last Modified: 2025-12-11 15:09:39+00:00
lidar/2024-11-12/casals_l1b_20241112T164340_001_02.h5
Size: 12.92 GB
Last Modified: 2025-12-11 15:09:39+00:00
lidar/2024-11-12/casals_l1b_20241112T164406_001_02.h5
Size: 13.02 GB
Last Modified: 2025-12-11 15:09:39+00:00
Alternatively, you can choose to use s3fs instead.
# Create filesystem object for anonymous access
fs = s3fs.S3FileSystem(anon=True)
# Bucket information
bucket_name = 'casals-data'
prefix = 'lidar/'
# List files in the bucket
print("Available CASALS data files:\n")
files = fs.ls(f'{bucket_name}/{prefix}')
for file_path in files[:20]: # Limit to first 20
print(f"{file_path}")
# Get file info for size
info = fs.info(file_path)
print(f" Size: {info['size'] / (1024**3):.2f} GB")
print(f" Last Modified: {info.get('LastModified', 'N/A')}\n")Available CASALS data files:
casals-data/lidar/
Size: 0.00 GB
Last Modified: N/A
casals-data/lidar/00README_CASALS_GE_Tours.txt
Size: 0.00 GB
Last Modified: 2026-01-20 18:35:40+00:00
casals-data/lidar/2024-11-12
Size: 0.00 GB
Last Modified: N/A
casals-data/lidar/2024-11-18
Size: 0.00 GB
Last Modified: N/A
casals-data/lidar/CASALS_2024-11-12_All_Amplitudes_Tour.m4v
Size: 7.50 GB
Last Modified: 2026-01-20 18:35:40+00:00
casals-data/lidar/CASALS_2024-11-12_All_Heights_Tour.m4v
Size: 7.50 GB
Last Modified: 2026-01-20 18:35:40+00:00
casals-data/lidar/CASALS_2024-11-18_All_Amplitudes_Tour.m4v
Size: 14.01 GB
Last Modified: 2026-01-20 18:35:40+00:00
casals-data/lidar/CASALS_2024-11-18_All_Heights_Tour.m4v
Size: 14.00 GB
Last Modified: 2026-01-20 18:35:40+00:00
2. Open dataset with lazy loading¶
Using xarray with h5netcdf engine allows us to access specific subsets without loading the entire ~14 GB file into memory.
%%time
# S3 url of the file to read
s3url = 's3://casals-data/lidar/2024-11-18/casals_l1b_20241118T172656_001_02.h5'
# Open dataset with lazy loading - doesn't load data into memory yet
# storage_options={'anon': True} enables anonymous (public) access
ds = xr.open_dataset(s3url, engine='h5netcdf', storage_options={'anon': True})
print("Dataset opened successfully! (data not yet loaded)")Dataset opened successfully! (data not yet loaded)
CPU times: user 7.19 s, sys: 3.87 s, total: 11.1 s
Wall time: 39.4 s
<timed exec>:6: UserWarning: The 'phony_dims' kwarg now defaults to 'access'. Previously 'phony_dims=None' would raise an error. For full netcdf equivalence please use phony_dims='sort'.
3. Inspect Dataset Structure¶
Let’s examine the L1B file structure and key variables without loading the full data.
# Inspect the overall structure
print("Dataset structure:")
print(ds)
print("\n" + "="*60)
print("Data variables (measurements and derived products):")
print("="*60)
for var in list(ds.data_vars):
print(f" {var}")
print("\n" + "="*60)
print("Dimensions:")
print("="*60)
for dim, size in dict(ds.dims).items():
print(f" {dim}: {size}")
print("\n" + "="*60)
print("Coordinates (indices for data organization):")
print("="*60)
for coord in list(ds.coords):
print(f" {coord}")Dataset structure:
<xarray.Dataset> Size: 21GB
Dimensions: (delta_time: 3604480, rx_bins: 2728,
phony_dim_0: 14080, phony_dim_1: 256,
tx_bins: 80)
Coordinates:
* delta_time (delta_time) float64 29MB 2.172e+08 ......
* rx_bins (rx_bins) int64 22kB 0 1 2 ... 2726 2727
* tx_bins (tx_bins) int64 640B 0 1 2 3 ... 77 78 79
Dimensions without coordinates: phony_dim_0, phony_dim_1
Data variables: (12/65)
bg_mean (delta_time) float64 29MB ...
bg_std (delta_time) float64 29MB ...
bin_size (delta_time) float64 29MB ...
dac (delta_time) float64 29MB ...
geoid (delta_time) float32 14MB ...
geoid_free2mean (delta_time) float32 14MB ...
... ...
tide_ocean (delta_time) float32 14MB ...
tide_ocean_pole (delta_time) float32 14MB ...
tide_pole (delta_time) float32 14MB ...
track_num (delta_time) int64 29MB ...
tracks (phony_dim_1) int64 2kB ...
tx_waveform (delta_time, tx_bins) int16 577MB ...
Attributes: (12/25)
ard_file: casals_ard_20241118T172656_001_02.h5
casals_epoch_gps_sec: 1198800018.0
casals_epoch_gps_sow: 86418.0
casals_epoch_gps_week: 1982
casals_epoch_utc_a: 2018-01-01t00:00:00.000000Z
end_delta_time: 217186041.14098355
... ...
speed_of_light: 299792458.0
start_delta_time: 217186016.11141512
start_utca: 2024-11-18T17:26:56.111415Z
style: 1D
tdms_file: casals_18-11-2024_12-27-36_151.tdms
ticks: 8
============================================================
Data variables (measurements and derived products):
============================================================
bg_mean
bg_std
bin_size
dac
geoid
geoid_free2mean
good_snr
instrument_altitude
instrument_altitude_error
instrument_latitude
instrument_latitude_error
instrument_longitude
instrument_longitude_error
local_beam_azimuth
local_beam_azimuth_error
local_beam_elevation
local_beam_elevation_error
range_bias_correction
refh
refh_amp
refh_bounce_time_offset
refh_bounce_time_offset_error
refh_error
refh_latitude
refh_latitude_error
refh_longitude
refh_longitude_error
refh_neutat_delay_derivative
refh_neutat_delay_total
refh_snr
refh_thres
rwstart
rwstart_bounce_time_offset
rwstart_bounce_time_offset_error
rwstart_error
rwstart_latitude
rwstart_latitude_error
rwstart_longitude
rwstart_longitude_error
rwstart_neutat_delay_derivative
rwstart_neutat_delay_total
rwstop
rwstop_bounce_time_offset
rwstop_bounce_time_offset_error
rwstop_error
rwstop_latitude
rwstop_latitude_error
rwstop_longitude
rwstop_longitude_error
rwstop_neutat_delay_derivative
rwstop_neutat_delay_total
rx_waveform
solar_azimuth
solar_elevation
sweep_num
sweeps
tide_earth
tide_earth_free2mean
tide_load
tide_ocean
tide_ocean_pole
tide_pole
track_num
tracks
tx_waveform
============================================================
Dimensions:
============================================================
delta_time: 3604480
rx_bins: 2728
phony_dim_0: 14080
phony_dim_1: 256
tx_bins: 80
============================================================
Coordinates (indices for data organization):
============================================================
delta_time
rx_bins
tx_bins
/tmp/ipykernel_127/3837883000.py:14: FutureWarning: The return type of `Dataset.dims` will be changed to return a set of dimension names in future, in order to be more consistent with `DataArray.dims`. To access a mapping from dimension names to lengths, please use `Dataset.sizes`.
for dim, size in dict(ds.dims).items():
4. Examine Key Variables¶
Important L1B datasets:
| Variable | Description |
|---|---|
tx_waveform | Transmit waveform data |
rx_waveform | Receive waveform data (contains surface returns) |
refh | Reference height - geolocated height of the waveform bin with maximum amplitude (WGS84 ellipsoidal height) |
tx_bins | List of Tx waveform bin indices |
rx_bins | List of Rx waveform bin indices |
delta_time | Laser transmit time in GPS seconds since 2018-01-01 |
sweep_num | Unique 0-based counter for each sweep |
track_num | Track (or channel) number |
# Check shapes of key waveform variables
if 'tx_waveform' in ds:
print(f"tx_waveform shape: {ds.tx_waveform.shape}")
print(f"tx_waveform dims: {ds.tx_waveform.dims}")
print(f" -> Dimensions: (pulses × bins)")
if 'rx_waveform' in ds:
print(f"\nrx_waveform shape: {ds.rx_waveform.shape}")
print(f"rx_waveform dims: {ds.rx_waveform.dims}")
print(f" -> Dimensions: (pulses × bins)")
if 'refh' in ds:
print(f"\nrefh (reference height) shape: {ds.refh.shape}")
print(f"refh dims: {ds.refh.dims}")
print(f" -> One reference height per pulse")tx_waveform shape: (3604480, 80)
tx_waveform dims: ('delta_time', 'tx_bins')
-> Dimensions: (pulses × bins)
rx_waveform shape: (3604480, 2728)
rx_waveform dims: ('delta_time', 'rx_bins')
-> Dimensions: (pulses × bins)
refh (reference height) shape: (3604480,)
refh dims: ('delta_time',)
-> One reference height per pulse
5. Efficient Data Subsetting¶
Strategy: Load only indexing variables first (lightweight), then use them to create masks for the data you actually need. This avoids loading the full ~14 GB file when you only need a small subset.
track_num = ds.track_num.load()
sweep_num = ds.sweep_num.load()
print(f"Loaded indexing variables:")
print(f" track_num: {len(track_num)} values (track/channel numbers)")
print(f" sweep_num: {len(sweep_num)} values (sweep numbers)")
print(f"\nTrack range: {track_num.min().values} to {track_num.max().values}")
print(f"Sweep range: {sweep_num.min().values} to {sweep_num.max().values}")Loaded indexing variables:
track_num: 3604480 values (track/channel numbers)
sweep_num: 3604480 values (sweep numbers)
Track range: 0 to 255
Sweep range: 0 to 14079
6. Select Data Subset¶
Selection criteria:
Tracks 0-255: All available wavelengths/channels
Sweeps 1000-1001: A small subset for visualization
This demonstrates how to efficiently extract specific pulses from the large dataset.
%%time
# Set the data selection parameters
track0 = 0 # Start track (channel)
track1 = 255 # End track (all channels)
sweep0 = 1000 # Start sweep
sweep1 = 1001 # End sweep (just 2 sweeps for this example)
# Create boolean mask to identify pulses matching our criteria
mask = ((track_num >= track0) & (track_num <= track1) &
(sweep_num >= sweep0) & (sweep_num <= sweep1))
# Get indices where mask is True
indices = np.where(mask)[0]
print(f"Found {len(indices)} matching pulses (records)")
print(f" Tracks: {track0} to {track1}")
print(f" Sweeps: {sweep0} to {sweep1}")
# Now load only the waveform data we need at those specific indices
# This is much faster than loading the entire dataset!
tx_waveform = ds.tx_waveform.isel(delta_time=indices).load()
rx_waveform = ds.rx_waveform.isel(delta_time=indices).load()
print(f"\nLoaded waveform data:")
print(f" tx_waveform: {tx_waveform.shape}")
print(f" rx_waveform: {rx_waveform.shape}")Found 512 matching pulses (records)
Tracks: 0 to 255
Sweeps: 1000 to 1001
Loaded waveform data:
tx_waveform: (512, 80)
rx_waveform: (512, 2728)
CPU times: user 1.17 s, sys: 469 ms, total: 1.64 s
Wall time: 8.09 s
7. Visualize TX (Transmit) Waveforms¶
TX waveforms show the outgoing laser pulse characteristics. Multiple waveforms are plotted to show variation across the selected pulses.
%%time
# Plot the Tx Waveforms
plt.figure(figsize=(12, 6))
for i in range(len(indices)):
plt.plot(ds.tx_bins, tx_waveform[i], alpha=0.7, linewidth=1)
plt.xlabel('TX Bin Number', fontsize=12)
plt.ylabel('Amplitude', fontsize=12)
plt.title(f'CASALS TX Waveforms (Sweeps {sweep0}-{sweep1}, Tracks {track0}-{track1})',
fontsize=14, fontweight='bold')
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
print(f"Plotted {len(indices)} TX waveforms")
Plotted 512 TX waveforms
CPU times: user 689 ms, sys: 3.67 ms, total: 692 ms
Wall time: 692 ms
8. Visualize RX (Receive) Waveforms¶
RX waveforms show the return signal including:
Surface returns (peaks in the waveform)
Atmospheric scattering
Background noise
%%time
# Plot the Rx Waveforms
plt.figure(figsize=(12, 6))
for i in range(len(indices)):
plt.plot(ds.rx_bins, rx_waveform[i], alpha=0.7, linewidth=1)
plt.xlabel('RX Bin Number', fontsize=12)
plt.ylabel('Amplitude', fontsize=12)
plt.title(f'CASALS RX Waveforms (Sweeps {sweep0}-{sweep1}, Tracks {track0}-{track1})',
fontsize=14, fontweight='bold')
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
print(f"Plotted {len(indices)} RX waveforms")
print("\nNote: The peaks in RX waveforms represent surface returns")
print("The bin with maximum amplitude is used to determine the reference height (refh)")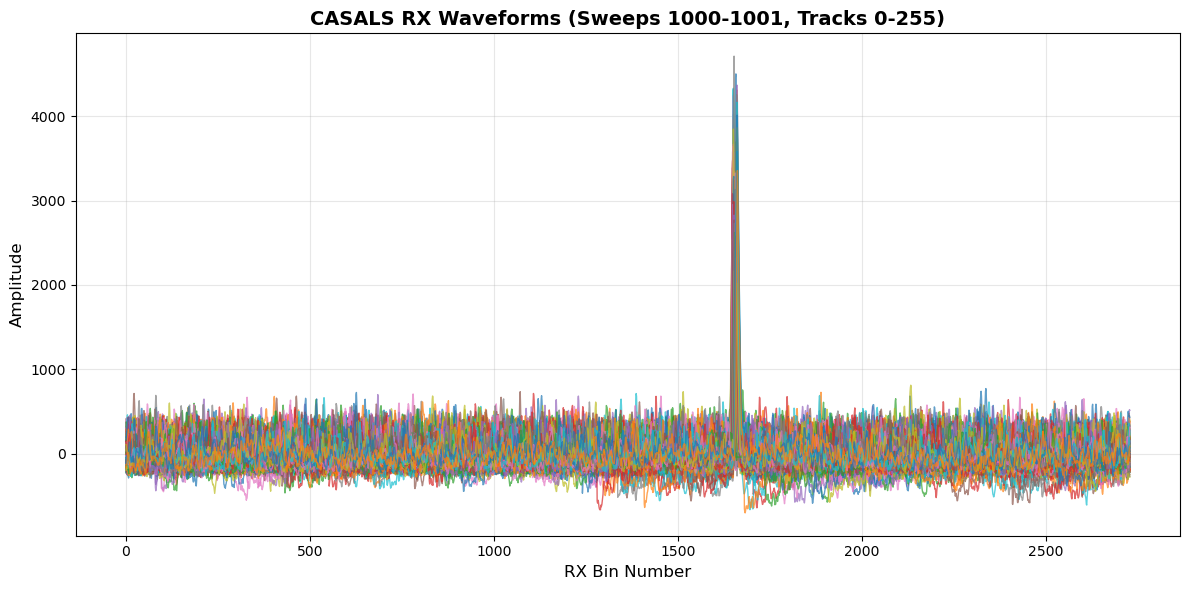
Plotted 512 RX waveforms
Note: The peaks in RX waveforms represent surface returns
The bin with maximum amplitude is used to determine the reference height (refh)
CPU times: user 2.15 s, sys: 34.2 ms, total: 2.19 s
Wall time: 2.19 s
9. Zoom into RX Waveform Peaks¶
Key L1B variable: refh (reference height)
Defined as: Geolocated height of the WF bin with the maximum amplitude (WGS84 ellipsoidal height)
The peaks we see here are the surface returns used to determine refh
Each pulse has associated uncertainty estimates (
refh_error)
%%time
# Zoom into the peak region to see surface returns in detail
peak0 = 1625 # Start bin for zoom
peak1 = 1700 # End bin for zoom
plt.figure(figsize=(12, 6))
for i in range(len(indices)):
plt.plot(ds.rx_bins[peak0:peak1], rx_waveform[i, peak0:peak1],
alpha=0.7, linewidth=1.5)
plt.xlabel('RX Bin Number', fontsize=12)
plt.ylabel('Amplitude', fontsize=12)
plt.title(f'CASALS RX Waveform Peaks - Surface Returns (bins {peak0}-{peak1})',
fontsize=14, fontweight='bold')
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
print(f"Zoomed view shows surface return peaks")
print(f"The bin with maximum amplitude in this region is used for reference height")
Zoomed view shows surface return peaks
The bin with maximum amplitude in this region is used for reference height
CPU times: user 728 ms, sys: 16.2 ms, total: 744 ms
Wall time: 743 ms
11. Summary of L1B CASALS data¶
L1B Global Metadata Attributes:¶
Names of input TDMS, L1A, GPS, ARD, and Geolocation files
Time epoch information (J2000 UTC-A format)
Number of Pulses, Sweeps, Tracks
Number of Tx and Rx Waveform Bins
Begin/End time of data
Processing constants (speed of light, conversions, etc.)
Key Dataset Categories:¶
1. Identifying Datasets (per pulse)¶
delta_time,sweep_num,track_num
2. Instrument Datasets (per pulse)¶
Instrument position and errors
3. Range Window Datasets (per pulse)¶
rwstart,rwstop(with position/errors)
4. Reference Height Datasets (per pulse)¶
refh(with position, amplitude, SNR, errors)
5. Beam Angle Datasets (per pulse)¶
local_beam_azimuth,local_beam_elevation(with errors)
6. Solar Angle Datasets (per pulse, at refh)¶
solar_azimuth,solar_elevation
7. Geophysical Datasets (per pulse, at refh)¶
Tide corrections, geoid, dac
8. Additional Quality Datasets¶
bg_mean,bg_std,bin_size,good_snrrange_bias_correction,refh_amp,refh_snr,refh_thres
12. Accessing Data from a Local Computer¶
If you’re running this notebook on your local machine (not in a cloud environment), you’ll need to configure AWS credentials for S3 access.
Three Access Methods
There are three different approaches:
Anonymous access (for public buckets like CASALS)
AWS credentials (for authenticated access)
Download locally first (for slower connections or repeated analysis)
Option 1: Use Anonymous (Public) Access¶
Since the data is in a public bucket, you can use anonymous access - same as in cloud environments!
import xarray as xr
# For public buckets - same as cloud environment
s3url = 's3://casals-data/lidar/2024-11-18/casals_l1b_20241118T172656_001_02.h5'
ds = xr.open_dataset(s3url, engine='h5netcdf', storage_options={'anon': True})Option 2: Configure AWS Credentials¶
For accessing data that requires authentication, you have several options:
Method 1: Using Environment Variables¶
# Set these in your terminal before running Python:
# export AWS_ACCESS_KEY_ID='your_access_key'
# export AWS_SECRET_ACCESS_KEY='your_secret_key'
# export AWS_DEFAULT_REGION='us-west-2'
import xarray as xr
s3url = 's3://casals-data/lidar/2024-11-18/casals_l1b_20241118T172656_001_02.h5'
ds = xr.open_dataset(s3url, engine='h5netcdf')Method 2: Using AWS CLI Configuration¶
# Run in terminal:
aws configure
# Then provide your credentials when prompted
# This stores credentials in ~/.aws/credentialsThen use the dataset normally in Python:
import xarray as xr
s3url = 's3://casals-data/lidar/2024-11-18/casals_l1b_20241118T172656_001_02.h5'
ds = xr.open_dataset(s3url, engine='h5netcdf')Method 3: Specify Credentials Directly¶
storage_opts = {
'key': 'your_access_key',
'secret': 'your_secret_key',
'client_kwargs': {'region_name': 'us-west-2'}
}
ds = xr.open_dataset(s3url, engine='h5netcdf', storage_options=storage_opts)Option 3: Download File Locally First¶
For slower internet connections or repeated analysis, download the file once:
import boto3
from botocore import UNSIGNED
from botocore.config import Config
import xarray as xr
# Configure S3 client
s3 = boto3.client('s3',
region_name='us-west-2',
config=Config(signature_version=UNSIGNED))
# Download file
bucket_name = 'casals-data'
file_key = 'lidar/2024-11-18/casals_l1b_20241118T172656_001_02.h5'
local_filename = '~/Downloads/casals_l1b_local.h5' # adjust this to fit where you want your file saved on your computer
print(f"Downloading {file_key}...")
print(f"This is a ~14 GB file, it may take several minutes...")
s3.download_file(bucket_name, file_key, local_filename)
print(f"Download complete! Saved as {local_filename}")
# Now open the local file
ds = xr.open_dataset(local_filename, engine='h5netcdf')
print("Dataset opened from local file")
# Use the dataset as normal
print(ds)Required Python Packages¶
Install these packages on your local machine:
Using mamba (recommended):¶
mamba install -c conda-forge xarray h5netcdf s3fs boto3 matplotlib numpyOr using pip:¶
pip install xarray h5netcdf s3fs boto3 matplotlib numpyTutorial Complete! 🎉¶
Happy analyzing! 🚀